Overview
Senior Investigator
Research Areas (IRP Lab Groups)
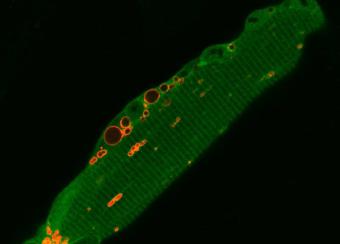
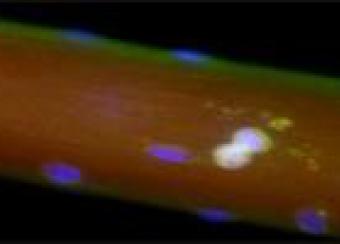
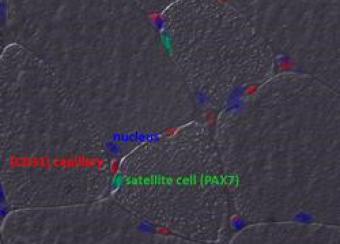
The Laboratory of Muscle Stem Cells & Gene Regulation investigates the epigenetics and metabolic changes in muscle stem cells, muscle-resident cells, and myofibers in adulthood and aging.
Our ultimate goal is to develop dietary and pharmacological interventions that target chromatin, transcriptional, and metabolic pathways in skeletal muscle cells. These interventions are designed to enhance muscle regeneration in muscular dystrophies and support the maintenance of muscle homeostasis and repair throughout adulthood and aging.
Areas of Interest
Specific areas of interest include:
- Transcriptional control of the regulatory circuits involved in various aspects of adult and aging muscle stem cell biology.
- The functional relationship between metabolites and epigenetic modifications impacting adult and aging muscle stem cell biology.
- The role of non-coding RNAs in skeletal muscle biology.
Core Research Facilities
Labs at the NIAMS are supported by the following state-of-the-art facilities and services:
Staff

Job Vacancy: Postdoctoral positions are immediately available in the Laboratory of Muscle Stem Cells and Gene Regulation (LMSCGR) at NIAMS at the National Institutes of Health, Bethesda, MD, USA. Ambitious and strongly motivated individuals with a solid background in molecular biology (bioinformatics is desirable but not required) are invited to apply.
Former Lab Members
- Morgan McKenzie, MD/PhD student, University of Colorado Anschutz
- Anthony Ascoli, MD, Resident Physician, PennMedicine, Philadephia, PA
- Giuseppina Caretti, PhD, Associate Professor, Department of Genetics, University of Milan, Italy
- Daphney Clermont, MD MPH, Pediatrician, The Johns Hopkins Hospital, MD
- Antonio Costanzo, MD, Head and Professor, Dermatology Clinical Unit, Humanitas University, Milan, Italy
- Veronica Ciuffoli, PhD, Research Scientist, Stanford University
- Stefania Dell’Orso, PhD, Staff Scientist, Director Office of Science and Technology, NIAMS
- Assia Derfoul, PhD, Biologist, Muscle Disease Unit, LMSCGR, NIAMS-IRP
- Monica Di Padova, PhD, Assistant Professor, Department of Experimental Medicine, University of L’Aquila, Italy
- Gabriella Fiore, PhD, Research Associate, Comparative Genomic Center, James Cook University, Townsville, Australia
- Marcella Fulco, MD, PhD, career transition to biomedical patenting
- Shanil Haugen, PhD, Assistant Director, US Food and Drug Administration
- Simona Iezzi, PhD, Staff Scientist, Regina Elena Cancer Center, Rome, Italy
- Aster Juan, PhD, Scientific Review Officer, CSR, NIH
- Mamduh Khateb, PhD, Research Fellow, NCI
- Raj Letchuman, Undergraduate Student, Yale University
- Christopher Lopez, MD/PhD Student, Stanford University
- Frederic Lluis Vinas, PhD, Professor, Stamcelinstituut Leuven (SCIL), KU Leuven, Leuven, Belgium
- Aissa Kaba, MTM, MedTech Scholar, UC Berkeley&UCSF
- Miroslav Koulnis, PhD, Scientist, Sony Biotechnology, Boston, MA
- Joseph G. Marx, PhD, F.B.I. Employee
- Kambiz Mousavi, PhD, Senior Scientist, Moderna Therapeutics, Cambridge, MA
- Alessandra Pasut, PhD, Postdoctoral Fellow, Center for Cancer Biology, Leuven, Belgium
- Jelena Perovanovic, PhD, Computational Biologist, Recursion, Salt Lake City, UT
- Valentina Proserpio, PhD, Department of Life Sciences and Systems Biology, University of Turin
- Madison Quig, BS, Graduate Student, University of Oxford, UK.
- Giulia Riparini, PhD, Senior Scientist, AstraZeneca
- James G. Ryall, PhD, Chief Scientific Officer, Vow Food, Sydney, Australia
- Kayoko Saso, PhD, Pharmacist
- R. Louis Schiltz, PhD, Biologist, Laboratory of Receptor Biology and Gene Expression, NCI
- Pei-Fang Tsai, Ph.D. Principal Scientist, Fate Therapeutics, San Diego, CA
- Karinna Vivanco, Graduate Student, UC Berkeley/UCSF Bioengineering PhD Program
- Stan Wang, MD PhD, CEO & Founder, Thymmune Therapeutics, Non-Resident Senior Fellow Council of Strategic Risks