Overview
Deputy Clinical Director
Research Areas (IRP Lab Groups)
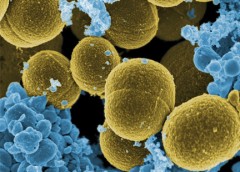
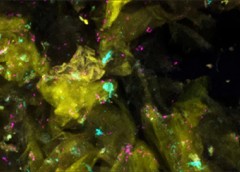
The Cutaneous Microbiome and Inflammation Section uses genomics to study the skin microbes in healthy individuals and patients with skin diseases and to expand our understanding of host-microbe interactions.
Our group studies the diversity and complexity of bacterial, fungal, and viral communities in healthy skin and in eczema from patients with atopic dermatitis and with primary immunodeficiencies. We integrate advanced genomic sequencing methods to investigate clinical samples and cultured isolates to more deeply understand these human microbial communities. Our work has demonstrated how the skin microbiome is patterned based on the location on the body surface (Grice et al., Science. 2009; Findley et al., Nature. 2013; Oh et al., Nature. 2014; Oh, Byrd et al., Cell. 2016), and can be distinguished in certain skin diseases and with specific immunodeficiencies (Kong et al., Genome Research. 2012; Byrd et al., Science Translational Medicine. 2017; Oh et al., Genome Research. 2013; Tirosh et al., Nature Medicine . 2018)
These studies have highlighted how the host shapes, and in turn may be shaped by, the skin microbiome. Our current efforts continue to explore these host-microbial interactions.
Research Focus
Our highly collaborative group studies the complexity of the microbial communities residing in and on the human body. We are focused on:

- Investigating the alterations in the human skin microbiome related to skin diseases, primary immunodeficiencies, and therapeutic interventions.
- Utilizing advances in microbiome and transcriptome sequencing to explore host-skin microbe interactions.
The translational team approach has been successful in integrating clinical medicine, genomics, microbiology, and immunology to study the role of staphylococci in atopic dermatitis as well as microbes in patients with immunodeficiencies. Our mission is to understand how microbes interact with the human host to elicit or ameliorate disease.
Related NIAMS Labs
Partner Labs at NIH
Lab Resources
Scientific Advances
Core Research Facilities
Labs at the NIAMS are supported by the following state-of-the-art facilities and services:
Staff

Our lab is always seeking to recruit the best talent in the scientific community. If you are interested in joining our team, please contact us.
Former Lab Members
Postdoctoral Fellows/Katz Scholars
- You Che, Ph.D., 2021-2024
- Jiwoong Choi, M.D., Ph.D., 2024-2025
- Jin Park, M.D., Ph.D., 2019-2021
- Hai Liang, Ph.D., 2018-2021
- Jay-Hyun Jo, Ph.D., 2015-2021
- Catriona P. Harkins, M.B.Ch.B., M.R.C.P., Ph.D., Katz Scholar in Dermatology Research, 2019-2020
- Martin Glatz, M.D., Visiting Fellow, 2012-2014
Medical Students
- Alexandra Firek, Medical Research Scholars Program Fellow, 2023-2024
- Margaret MacGibeny, Clinical Electives Program Student, 2020
- Therese Woodring, Medical Research Scholars Program Fellow, 2016-2017
- Radhika Nakrani, Medical Research Scholars Program Fellow, 2013-2014
Postbaccalaureate Fellows
- Emily Tung, 2024-2026
- Ryleigh Griffin, 2023-2025
- Milan Stolpman, 2022-2024
- Grace Suh, 2022-2024
- Coralys Citron, 2023-2023
- Layne Oram, 2021-2022
- Jessica Portillo, 2019-2021
- Meridith Pensler, 2020-2021
- Nicole Schwardt, 2017-2019
- Elizabeth A. Kennedy, 2015-2017
- Amanda Johnson, 2013-2015
- Elizabeth Bassett, 2007-2008
Image & Media Gallery
Clinical Trials
Atopic dermatitis, or eczema, is a chronic skin disorder. The use of antibiotics has revolutionized medicine, yet the impact of antimicrobials on the human microbiome is incompletely understood. This study aims to characterize microbiome alterations in healthy adult volunteers and patients with atopic dermatitis after antimicrobial treatments.
Skin diseases represent one of the most common medical problems in the United States, affecting 1 in 3 people at any given time. This study aims to procure biologic samples for exploratory cellular, molecular, genetic and genomic biological studies from subjects with dermatologic conditions, subjects at risk for developing dermatologic conditions and healthy volunteers in the support of NIH biomedical studies.