Overview
Senior Investigator
Research Areas (IRP Lab Groups)
The NIAMS Protein Expression Laboratory (PEL) studies the structure and function of viral proteins, primarily those of the Human Immunodeficiency Virus (HIV-1) and the Hepatitis B Virus (HBV). The PEL also studies the interactions of these viral proteins with host proteins.
Most structural biology techniques, especially those for studying the three-dimensional structures of proteins, require large quantities of highly purified, monodisperse, and correctly folded proteins. The PEL uses recombinant DNA techniques to produce proteins both for in-house use and for collaborating scientists. It employs techniques such as analytical ultracentrifugation, UV-CD, NMR, X-ray crystallography, and cryo-electron microscopy to characterize and determine the structures of these proteins and their complexes.
HIV-1:
gp41 and gp120: HIV-1 entry into cells is initiated when a trimeric complex on the virion surface, composed of gp41 and gp120, interacts with the principal cellular receptor, CD4. The trimeric complex is therefore a target for vaccine and drug development. When gp120 binds to the CD4 receptor it changes the conformation of gp41, which induces fusion between the viral and host membranes. Using both NMR and X-ray crystallography, the PEL and its collaborators have determined the structure of gp41. Based on this structural and biochemical information, a mechanism for membrane fusion has been proposed.
Protease: Early in HIV-1 infection, viral RNA is translated into a polyprotein which is then cleaved by the viral protease at nine sites to create the functional proteins. In this process, the dimeric protease catalyzes its own production. Without the protease, viral replication is not possible and so it is an important drug target. The virus evolves to evade these drugs, lessening their efficacy. Based on the study of protease residues susceptible to the oxidative process, namely the sulfur-containing residues methionine and cysteine, the PEL and its collaborators have proposed that drugs that target the region between the subunits may provide an alternate site of intervention.

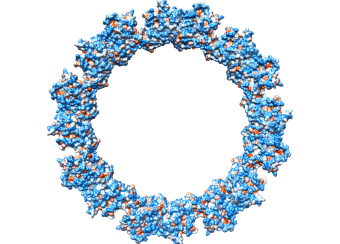
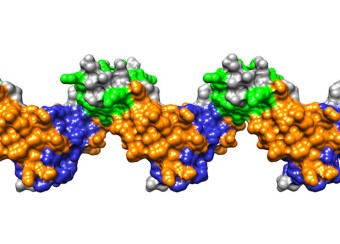
Rev: The Rev protein is an essential viral regulator that assembles on a highly structured 350-nucleotide motif, the Rev Response Element (RRE), in un-spliced and partially spliced viral transcripts, enabling their nuclear export. Despite over 30 years of active research, the molecular details whereby Rev binds to and assembles on the RRE remain largely unknown. Structural details of this complex nucleoprotein assembly would open up new drug targets. A model of the RRE, with a Rev tetramer docked onto it in accordance with X-ray crystallography, cryo-EM, NMR, SAXS, SHAPE, and mutational analyses, has been developed by the PEL ( see Fig. 1 ). The structure of an RNA aptamer that can inhibit HIV-1 by blocking Rev-RRE binding and Rev-Rev association has been determined. Engineered antibodies, their fragments (Fabs), and cyclic peptides based on the antibody CDR loops, that inhibit HIV-1 have been devised. A patent application has been filed. Rev also interacts with several host proteins which may present therapeutic targets. These include the nuclear regulatory factors B23 and Nap1, and tubulin, which are all being studied by the PEL.

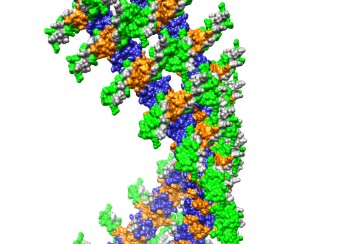
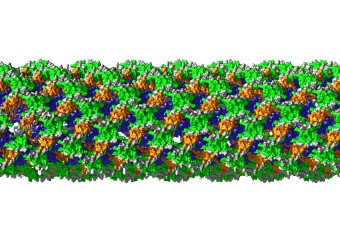
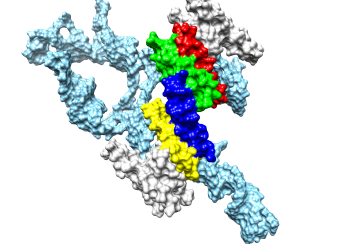
Rev and Tubulin: The PEL has found that Rev interacts strongly and stoichiometrically with tubulin to form double-ring complexes in vitro. The PEL has also shown that treatment of tubulin with Cryptophycin-1 (a compound derived from marine cyanobacteria of the genus Nostoc with anticancer activity 1000-fold greater than paclitaxel) forms highly stable rings composed of eight tubulin dimers and that Rev binds to these to also form double-rings. The binding sites on tubulin, and the mechanisms of ring formation, are not understood for either Cryptophycin-1 or Rev. The PEL is studying the structure of these complexes by cryo-electron microscopy using the Krios microscope at the NIH Multi Institute Cryo-EM Facility (MICEF) ( see Fig. 2 ). The PEL has now also formed complexes of Hela cell tubulin with Cryptophycin-52, an analog of Cryptophycin-1 that has previously advanced to phase II clinical trials as a chemotherapeutic. This will allow us to understand the mechanisms of these anticancer drugs and how they can be improved, something of longstanding interest to both the tubulin and cancer research communities.
Hepatitis B Virus:
HBV core-antigen: HBV infects approximately one-third of the human population, is the major cause of liver cancer worldwide and causes almost 1 million deaths annually. Although a vaccine has been developed, it is not universally available and, as the virus is vertically transmitted from mothers to infants, HBV is often acquired in childhood and becomes a chronic condition.

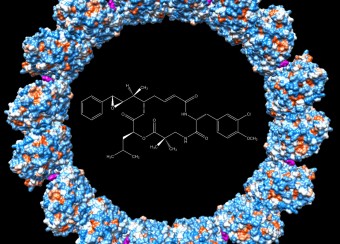
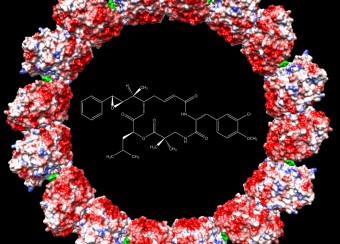
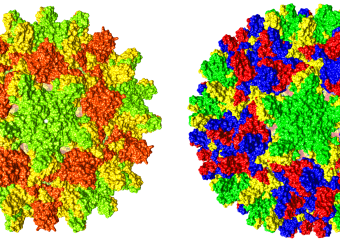
The viral capsid plays an important structural and metabolic role in the replication cycle of any virus and, being distinct from host components, is a potential target for intervention. In the case of HBV, the dimeric capsid protein assembles into shells of two sizes, called T=3 and T=4 in reference to their symmetry ( see Fig. 3 ). The PEL studies the assembly and structure of these capsids using recombinant proteins, and by making extensive use of mutagenesis and structural analysis by X-ray crystallography and electron microscopy. The PEL also studies the binding to capsids of HBV-patient and monoclonal antibodies and it has engineered antibody fragments (Fab and scFv) that effectively block capsid assembly.
HBV e-antigen: The viral e-antigen is a dimeric protein that subverts the human immune system to establish a chronic infection. The mechanism is not known. The e-antigen polypeptide is essentially identical to that of the core-antigen except that it has a 10-residue propeptide. The PEL has found that this difference causes the dimer to adopt a radically different arrangement from that of core-antigen and that it can be converted to an arrangement like the latter by reduction.
The PEL has produced a panel of chimeric (human framework) anti-e-antigen rabbit antibodies (Fabs), some with very high affinities (Kd = 10-12 M). These antibodies have both diagnostic and therapeutic potential. The PEL has developed a quantitative assay for the e-antigen which is superior to existing commercial assays both in terms of sensitivity and specificity. Using these antibodies, the PEL has immunoaffinity-purified e-antigen from individual patient plasmas and subjected it to proteomic analysis, demonstrating previously unknown heterogeneity in this antigen. The X-ray crystallographic structures of several of the chimeric antibodies (as scFv) complexed with e-antigen have been determined. These antibodies bind to epitopes forming parts of the dimer-dimer interface which mediates capsid assembly in the core-antigen (and reduced e-antigen). This and future work will provide a structural framework to test the strategy of treating chronic HBV using antibodies (and derived mimics) that target both the e-antigen and capsid assembly. A patent application has been filed. The nature of these antibodies (recombinant source, human framework, exceptionally high affinity, and specificity) also raises the potential for their use as antibody-drug complexes (ADCs) in the treatment of HBV infection.
Core Research Facilities
Labs at the NIAMS are supported by the following state-of-the-art facilities and services: